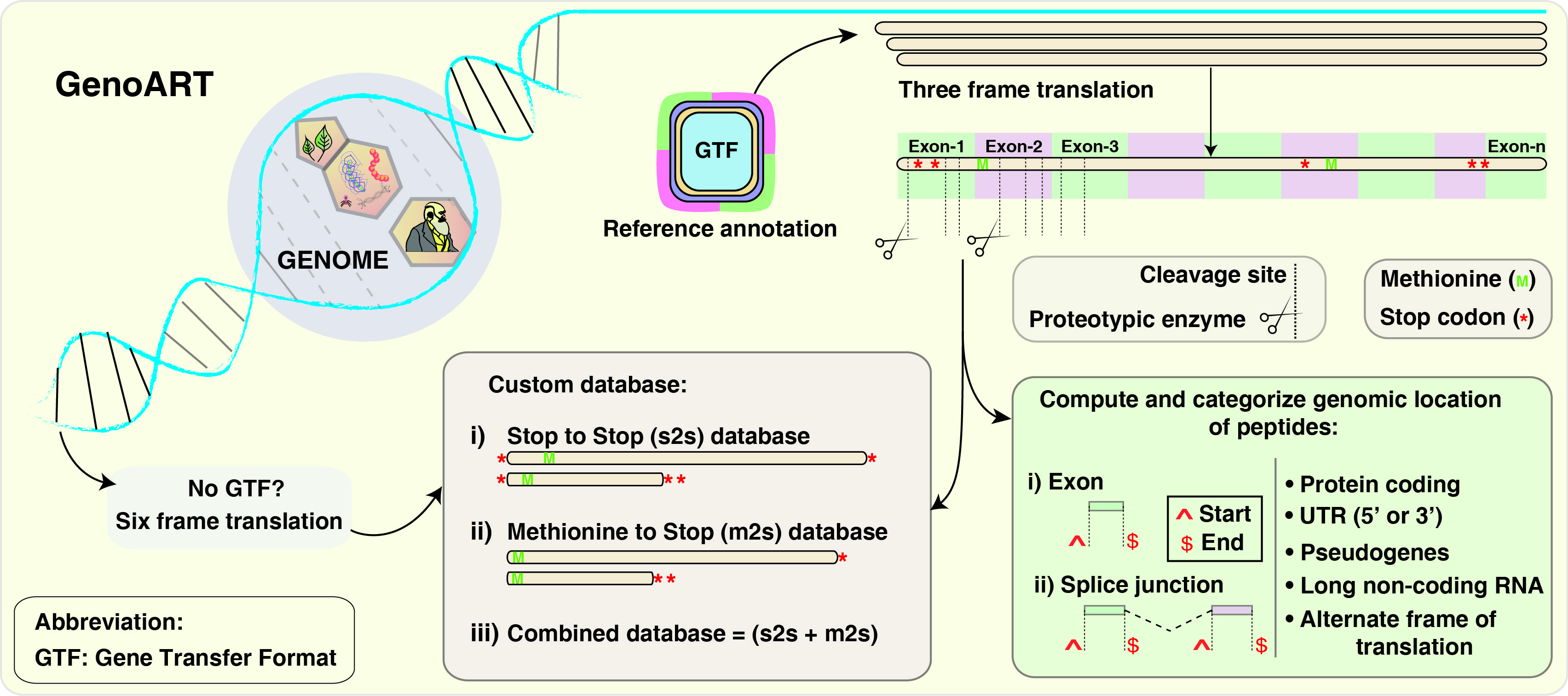
GenoART is a standalone application for detecting coding potential of transcripts complemented by experimentally derived proteomics (MS/MS) data. GenoART will make use of Python, PERL and javascripts for processing your input genome and create a custom database for proteogenomic searches. A detailed protocol is available online. In simple terms, the custom database generated by this application can be used as a reference protein database and search the unmatched/unassigned spectra in any search engines such as Mascot, SEQUEST, X! Tandem or Andromeda. GenoART can be queried to categorize and browse these resultant peptides at the click of a button. The peptides will include the genomic coordinates which allows user to navigate to UCSC genome browser which can facilitate more details about the gene/transcript and can be compaired with predicted genes, Expressed sequence Tag (EST), and perform comparitive genomics for more evidence. A GTF (Gene Transfer Format) will be available to download upon querying, which can later be viewed in any genome visualizer such as UCSC or IGV to compare accross different studies.
| Operating System | Download | Size (MB) | Remark (Click here to download sample dataset) |
|---|---|---|---|
| MS Windows | GenoART v1.0 - Win | 18.33 | Graphical user interface using Electron for MS Windows. |
| Linux | GenoART v1.0 - Linux | 05.08 | Graphical user interface using Electron for Linux. |
| Mac OS | GenoART v1.0 - Mac | 04.93 | Graphical user interface using Electron for Mac OS. |